This doesn't look sustainable
Breaking up is hard to do.
That’s certainly true for common plastics like polyethylene terephthalate (PET). A water bottle made of a thin film of PET (perhaps half a millimeter thick) takes about 450 years to degrade. Along the way it will exist as microplastics, which are so pervasive they are even turning up in living people’s lung tissue, as we saw for the first time just a month ago. And even those kinds of numbers are guesstimates because many studies don’t last long enough to see any appreciable degradation of PET at all.
A lot of efforts to produce biodegradable and bioresorbable plastics are making good progress, and that’s great for the future, but what about the mountains of plastic that already exist and that we keep on generating? In the U.S., the landfill rate for discarded plastics is still about 75%! We have a lot of work to do.
Good thing Hal Alper’s chemical engineering lab at the University of Texas is on the job. In the April 27 issue of Nature, they report on an enzyme they developed called FAST-PETase. Designed with the help of artificial intelligence, it degrades untreated postconsumer PET not in centuries, but in days. And this can be done at temperatures of 50°C and below, where many types of bacteria can thrive. See where this is going?
Before we delve into how this was done, let’s check out some results. Here is a plastic article that would normally take centuries to degrade, treated with FAST-PETase for the amount of time shown:
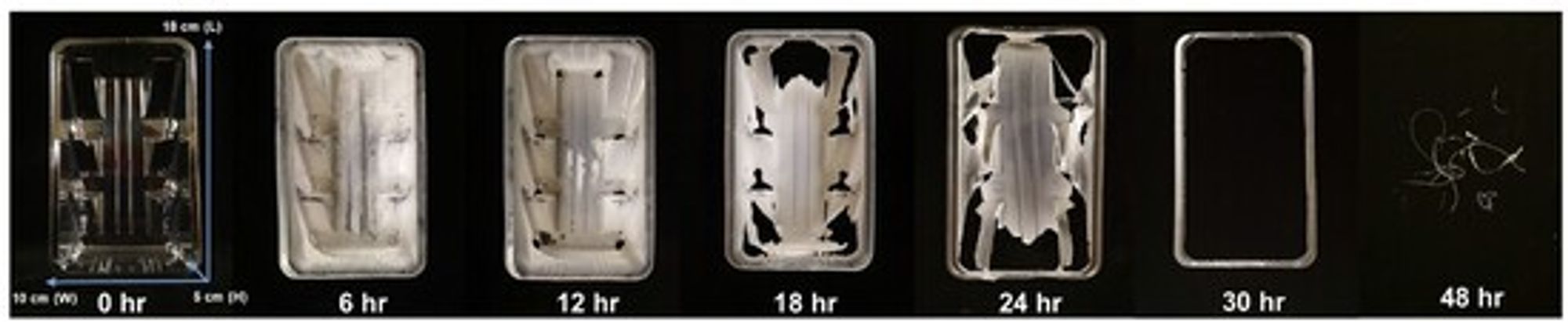
A plastic sardine can? Phone case? Whatever it is, it degrades completely in 48 hours
If you can’t make out the labels, those are 0, 6, 12, 18, 24, 30, and 48 hours. That thing is gone, not in centuries but in forty-eight hours. Come on, man! I’ve had cheese-induced stupors that have lasted longer than that.
Lest you think this particular plastic article was special in some way, the researchers cut out discs from a wide variety of untreated plastic consumer products and set the FAST-PETase enzyme to work on them. The top bar graph shows the number of days it took each disc to be degraded completely. The top number on the vertical axis is 8 days, meaning all those discs were gone within a week:
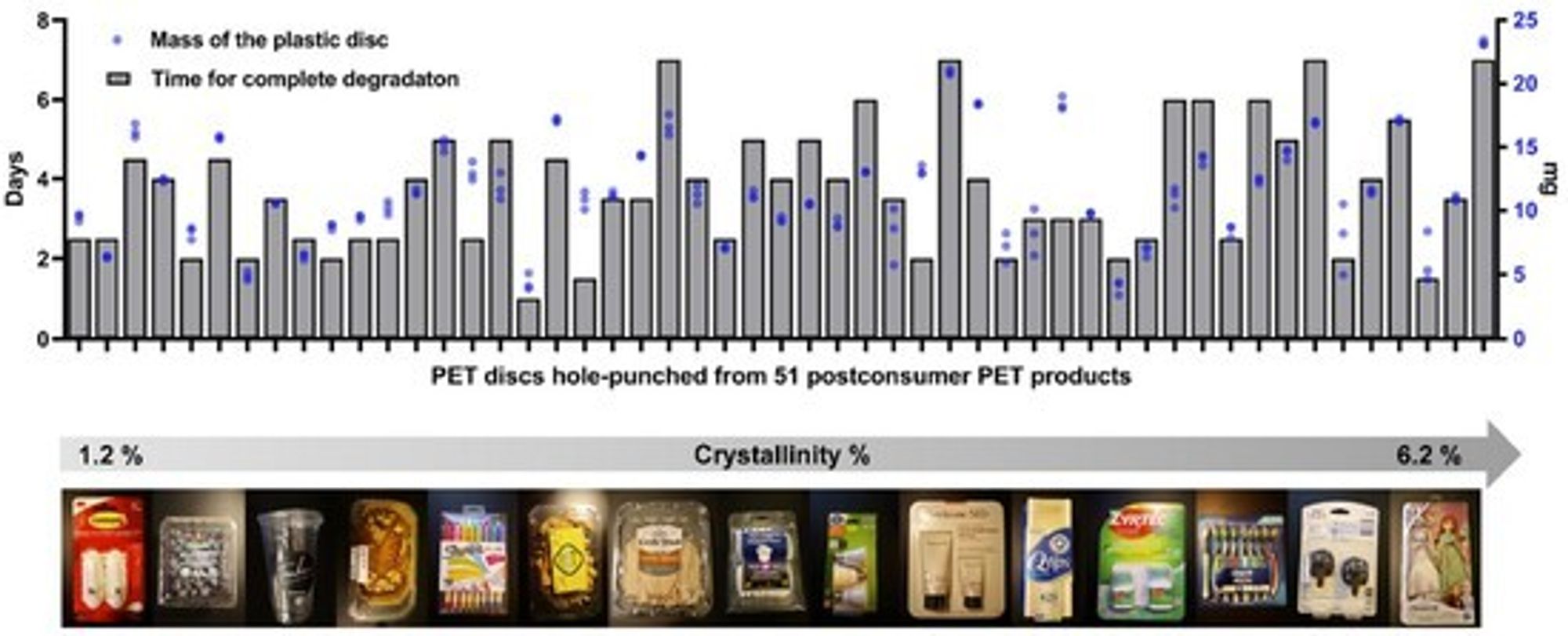
PET discs from 51 untreated postconsumer products are all completely degraded within a week at 50°C. Crystallinity (which can make plastic harder to degrade because of the tight attraction among polymer chains) had little or no effect here. Everything got devoured!
Here we have a variety of untreated plastic articles from the trash, over a range of crystallinities, disappearing in one week or less, with the help of biology.
How the heck did this happen?
There are bacteria out there that can eat plastic as their main source of carbon and energy. Not super quickly or efficiently, but they can do it. They became known to us humans in 2016, when we named them Ideonella sakaiensis. They were isolated outside a plastic bottle-recycling facility in Japan, and in that environment, they had evolved the capability to eat plastic.
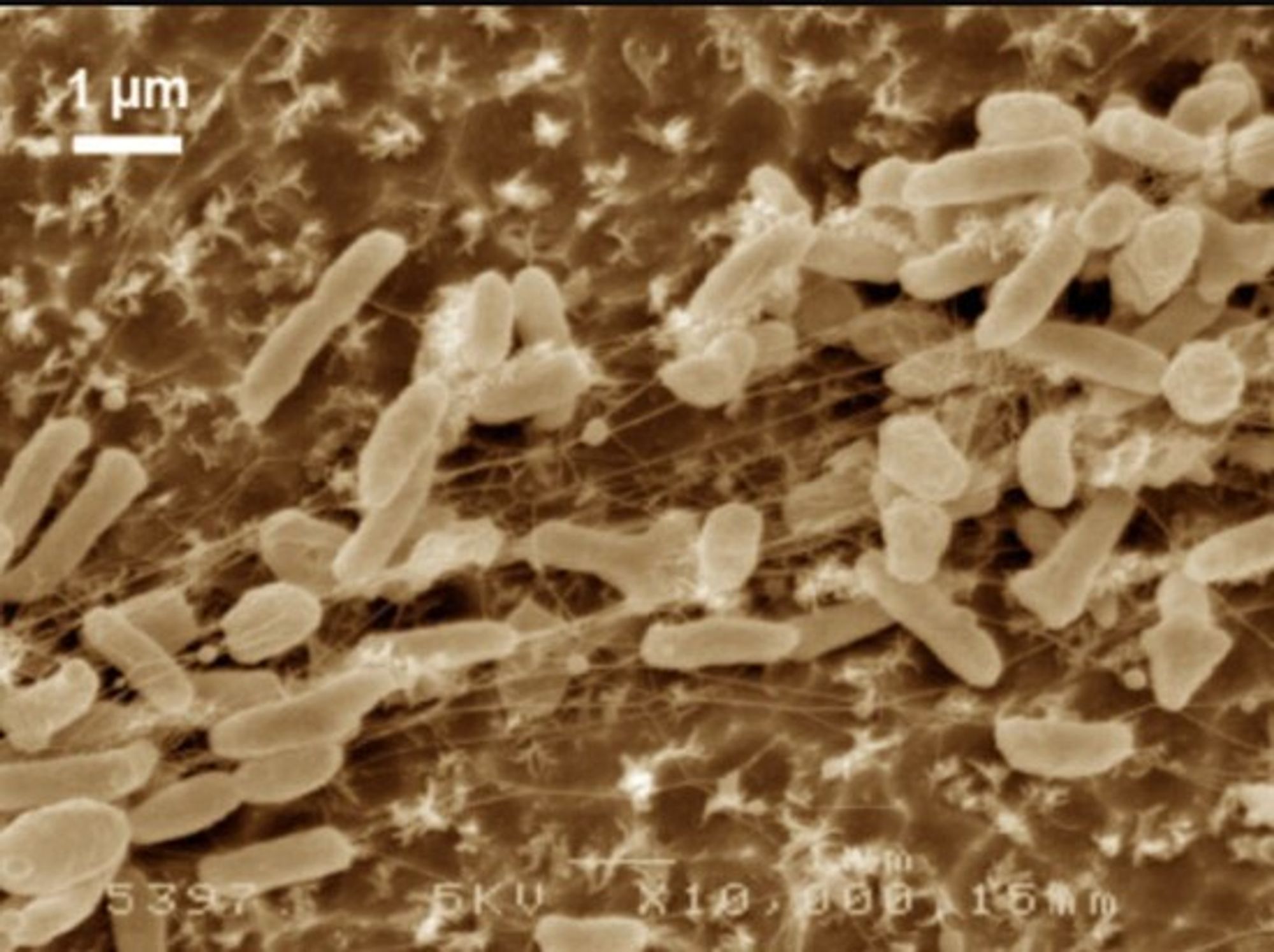
The plastic-eating bacterium Ideonella sakaiensis
They break the PET plastic down in two steps:
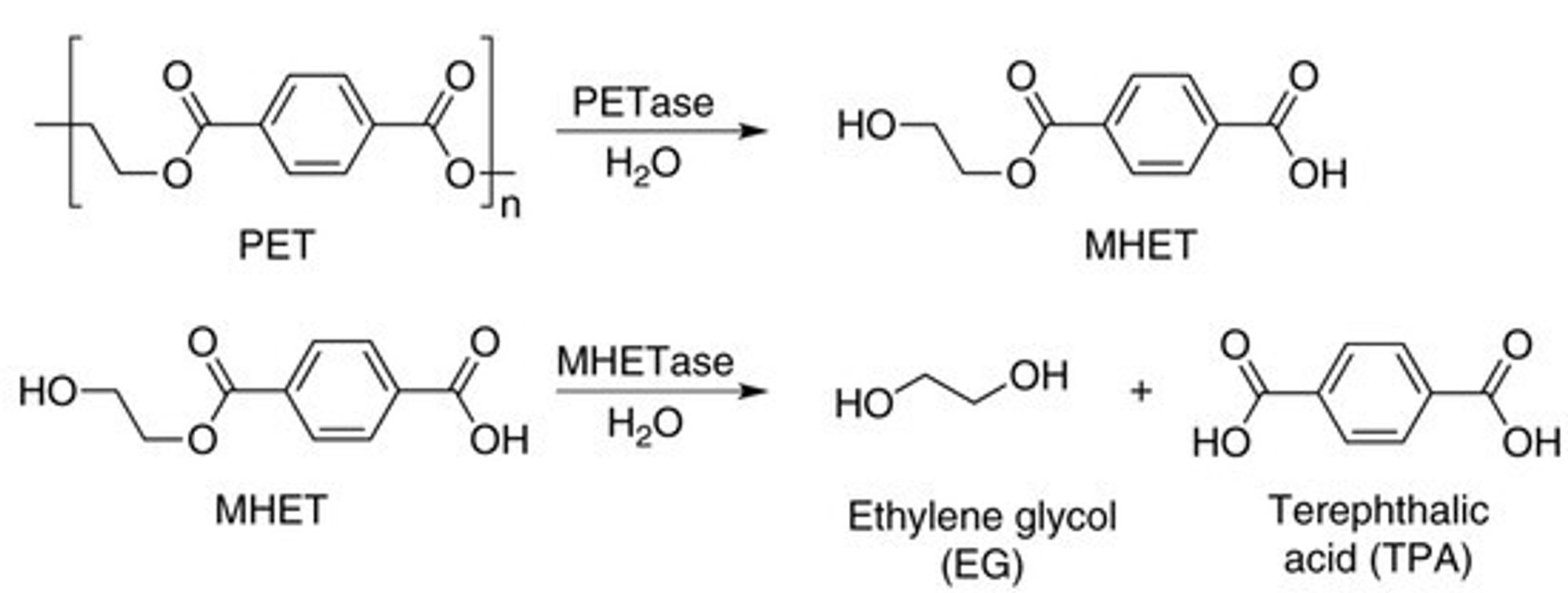
PET is a long chain of repeated units, one of which is shown in brackets. The subscript “n” means that a polymer can have any number (“n”) of these units, connected in a linear chain. For a useful plastic, “n” might be in the hundreds or thousands.
The difficult step is the first one, PETase, because these bacteria need to make and secrete an enzyme that can pry its way into the plastic and snip the C-O bonds that hold the polymer together. Once that happens, you end up with small molecules — MHET and ultimately ethylene glycol and terephthalic acid — and at that point any number of bacteria out there can easily chow down on these and convert them to CO2 and water. So we need to make PETase as efficient as we can.
PETase has a couple of notable limitations:
1) It isn’t very stable, losing all of its activity within 24 hours at 37°C 2) Its activity is only reasonably good at high temperatures (above 70°C)
So any process that hopes to use this enzyme at ambient-like temperatures is going to suffer from instability and low rates of activity. But, as in any evolutionary process, all we need is a foothold. We’ve got an enzyme here that can do what we want; we just need to make it more active and stable.
Fortunately, by 2018 the structure of PETase had been solved, so now it was known how this protein folds up and where all of its amino acids are located relative to one another.
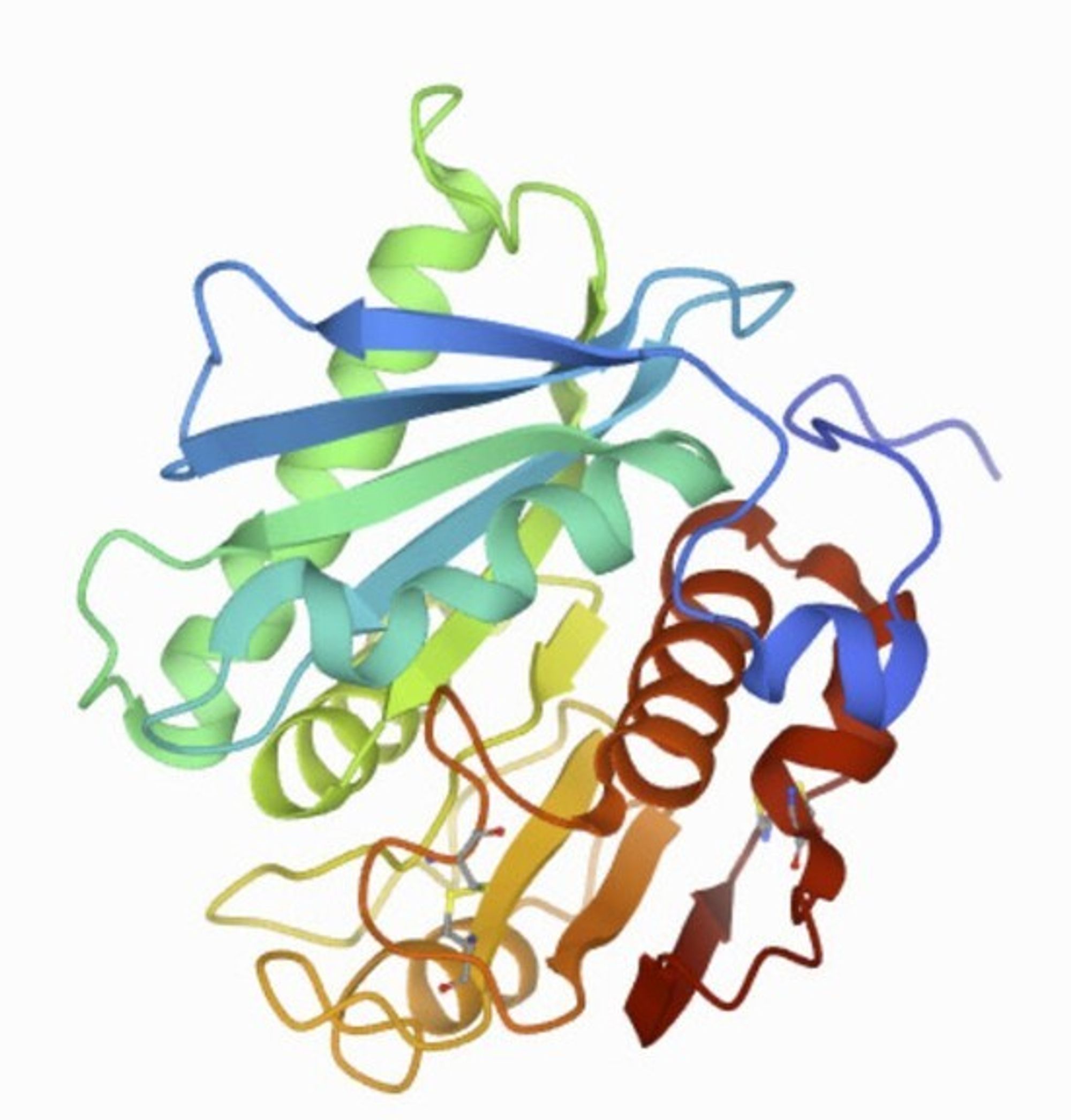
The structure of PETase from Ideonella sakaiensis
So a few groups started trying to modify PETase by substituting different amino acids at certain positions based on what they knew about that structure, but with only modest success. They got a little bit more stability, but the enzyme was still active enough to be practically useful only at high temperatures.
Another cool development that happened over the last few years is MutCompute, an artificial intelligence-based platform that can review any enzyme and judge how each of its amino acids fits into its local environment. This kind of review of PETase led to some very specific recommendations for making the enzyme more stable.
Of course, none of us here is a protein chemist (OK, maybe a couple of you are) but we can still look at one of the changes that MutCompute recommended so we can grasp visually why it might help:
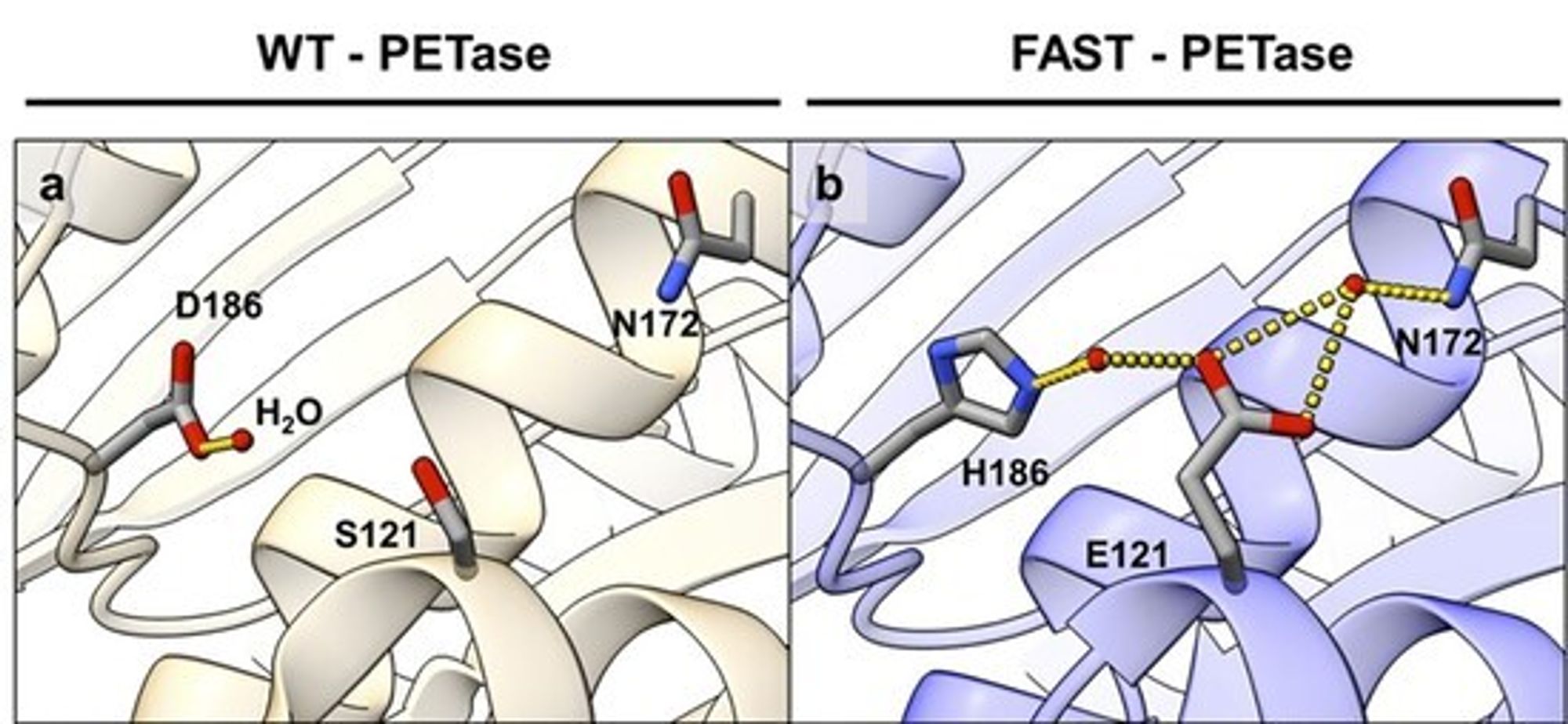
At left is the “wild-type” PETase isolated directly from the natural organism; at right is FAST-PETase. Here two simultaneous amino-acid changes make the hydrogen-bonding (yellow dashed lines) situation much different
We can simultaneously change two amino acids that are near each other in the protein to help them interact so they help hold the enzyme together, but we would never, ever guess these combinations without the help of computation that considers the entire structure of the enzyme.
If we change amino acid D186 (aspartic acid) at left to H186 (histidine) at right, and we change S121 (serine) at left to E121 (glutamic acid) at right, we get a microenvironment where some nice hydrogen bonds (shown by yellow dashed lines) can form in space among these amino acids to stabilize the overall structure in that little area, and yet we don’t change the structure of the overall enzyme too much. All the predicted twisties and curlicues still look pretty similar in the tan picture at left and the blue picture at right, but the bonds shown by dashed yellow lines look a lot different.
Now if you combine a few of these manipulations, it turns out you do indeed get a much more stable enzyme, one that can hold together and keep its activity for a much longer time, even at higher temperatures.
But what surprised me about this — and I’m sure it surprised the researchers too — is that this holistically engineered enzyme is also much more active at lower temperatures. Instead of having to crank the temperature up to 72°C (162°F), you get terrific activity at 50°C (122°F). In fact, the engineered FAST-PETase is more active at 50°C than any variant has ever been at 72°C. Chemical reactions go faster at higher temperatures because molecules are simply moving around faster, and so a reaction at 72°C should go about 25% faster than one at 50°C. Yet this new enzyme flips that on its head because its structure makes it that much more awesomely effective as a catalyst.
Most microorganisms are dead in the water at 72°C, but there are lots of bacteria and fungi that can thrive at 50°C. So at this point it comes down to putting the DNA encoding this enzyme (along with MHETase) into a friendly microorganism that does well at 50°C (a “thermophile”), and letting it feast on PET waste. We’ve become pretty good at getting our favorite DNA into thermophiles, so this is very doable.

Thermophilic organisms thrive in places like this hot spring in Yellowstone National Park
OR! Because this enzyme is now so stable, it could be mass-produced by some helpful bacterium we engineer it into (even good old E. coli), and THAT could be used in a mildly heated process to put landfill waste out to pasture in short order. We do this already in our laundry, where enzymes mass-produced in giant fermenters by bacteria break down the proteins, fats, and sugars we blop onto our clothes. It hadn’t been very possible with plastics like PET before, but with this development, suddenly it is.
This means that rapid disappearance of plastic waste could very well be coming to a landfill — or lake or ocean — near you.
If you’re concerned about the release of CO2 by such a process, that’s justified, but there’s more to this story!
Not only can this process be used to degrade PET, but in fact to reconstitute it. Once you break the polymer down into its components, you can let bacteria in the wild reduce them to CO2 and water. OR you can re-polymerize them, which is a nifty way of casting aside any impurities like dyes or product residue and regenerating “virgin” PET. This is a recycling program where you don’t have to worry too much about the state of the discarded plastic.
The authors walk us through this process pictorially, because they demonstrated that it works. Here they take green-dyed postconsumer PET, degrade it with FAST-PETase, recover the polymer building blocks, and repolymerize it into a colorless, effectively new product:
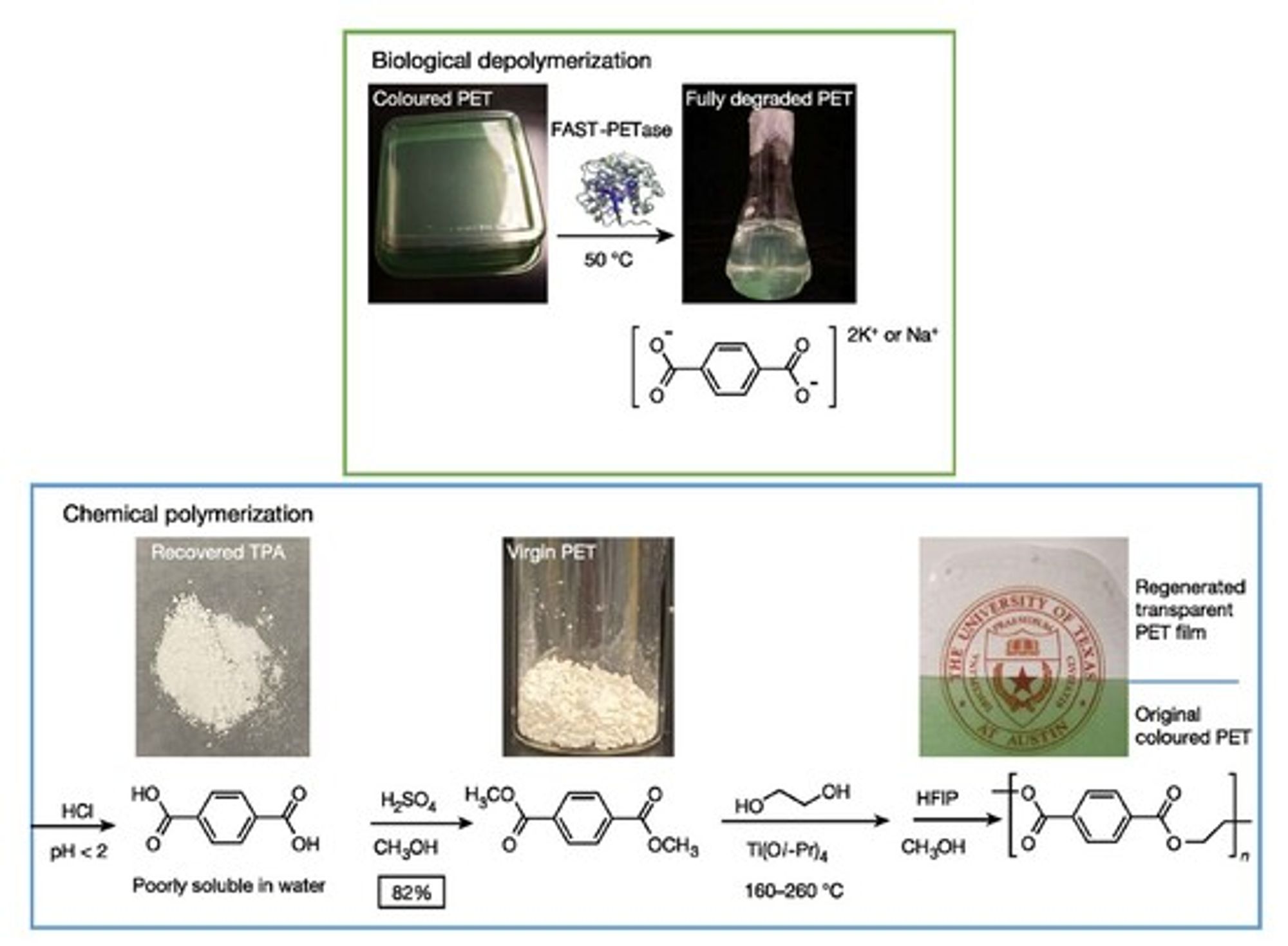
Regeneration of colorless PET from untreated green-dyed postconsumer PET
So while there is a lot of promise in biodegradable and compostable plastics, we aren’t going cold turkey on petroleum-based plastics anytime soon, and we aleady have mountains of plastic to deal with. But now we really are a step closer to actually being able to deal with them.
Bacteria, when faced with the need to survive, can become really smart all of a sudden. And when people have serious problems to contend with, if we keep our heads on straight and use reason to guide us, we can be really smart, too. This research drew from microbiology, organic chemistry, polymer science, chemical engineering, mathematics, and computer science. If a kid ever asks, Why do I have to learn that? When will I ever use that? (as my kids sometimes do), an accomplishment like this provides a good answer: Because you’ll be capable of more.
This new line of attack on plastic waste ultimately originated in Japan, so I’ll let Guitar Wolf close it out….. Fujiyama Attack!
x
Thanks very much to Hal Alper at the University of Texas for kindly sending me a reprint of the paper, and I hope I have done it justice here.