Fully assembled SEND packages are released from the cell to be collected for gene therapy. Credit: McGovern Institute
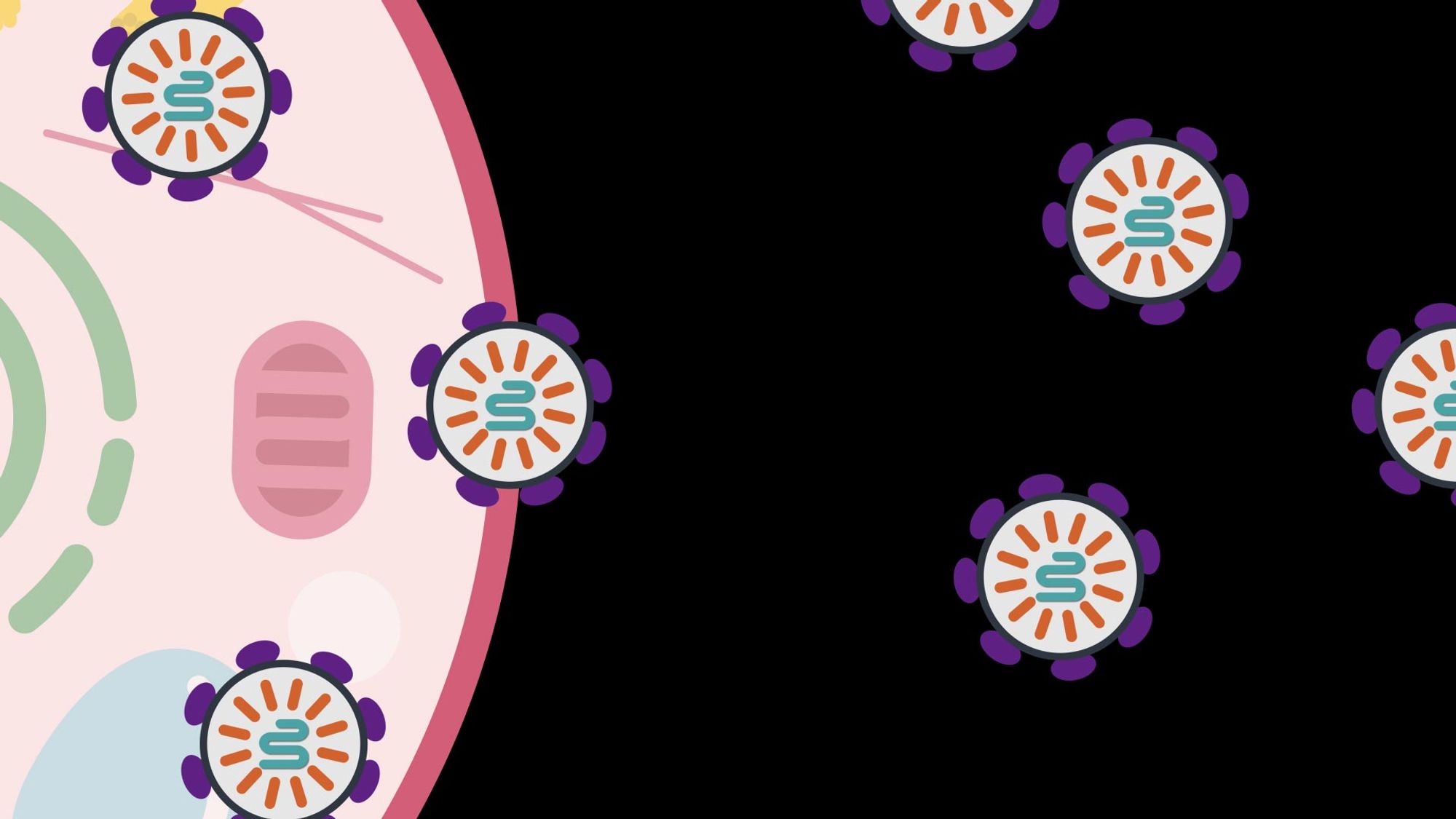
Made of components found in the human body, the programmable system is a step toward safer, targeted delivery of gene editing and other molecular therapeutics.
Researchers from MIT, the McGovern Institute for Brain Research at MIT, the Howard Hughes Medical Institute, and the Broad Institute of MIT and Harvard have developed a new way to deliver molecular therapies to cells. The system, called SEND, can be programmed to encapsulate and deliver different RNA cargoes. SEND harnesses natural proteins in the body that form virus-like particles and bind RNA, and it may provoke less of an immune response than other delivery approaches.
The new delivery platform works efficiently in cell models, and, with further development, could open up a new class of delivery methods for a wide range of molecular medicines — including those for gene editing and gene replacement. Existing delivery vehicles for these therapeutics can be inefficient and randomly integrate into the genome of cells, and some can stimulate unwanted immune reactions. SEND has the promise to overcome these limitations, which could open up new opportunities to deploy molecular medicine.
Researchers from MIT, the McGovern Institute for Brain Research at MIT, the Howard Hughes Medical Institute, and the Broad Institute of MIT and Harvard have developed a new way to deliver molecular therapies to cells. Credit: Produced by the McGovern Institute and Opus Design in collaboration with Feng Zhang, Rhiannon Macrae, and the Broad Institute.
“The biomedical community has been developing powerful molecular therapeutics, but delivering them to cells in a precise and efficient way is challenging,” said CRISPR pioneer Feng Zhang, senior author on the study, core institute member at the Broad Institute, investigator at the McGovern Institute, and the James and Patricia Poitras Professor of Neuroscience at MIT. “SEND has the potential to overcome these challenges.” Zhang is also an investigator at the Howard Hughes Medical Institute and a professor in MIT’s Departments of Brain and Cognitive Sciences and Biological Engineering.
Reporting in Science, the team describes how SEND (Selective Endogenous eNcapsidation for cellular Delivery) takes advantage of molecules made by human cells. At the center of SEND is a protein called PEG10, which normally binds to its own mRNA and forms a spherical protective capsule around it. In their study, the team engineered PEG10 to selectively package and deliver other RNA. The scientists used SEND to deliver the CRISPR-Cas9 gene editing system to mouse and human cells to edit targeted genes.
First author Michael Segel, a postdoctoral researcher in Zhang’s lab, and Blake Lash, second author and a graduate student in the lab, said PEG10 is not unique in its ability to transfer RNA. “That’s what’s so exciting,” said Segel. “This study shows that there are probably other RNA transfer systems in the human body that can also be harnessed for therapeutic purposes. It also raises some really fascinating questions about what the natural roles of these proteins might be.”
Inspiration from within
The PEG10 protein exists naturally in humans and is derived from a “retrotransposon” — a virus-like genetic element — that integrated itself into the genome of human ancestors millions of years ago. Over time, PEG10 has been co-opted by the body to become part of the repertoire of proteins important for life.
Four years ago, researchers showed that another retrotransposon-derived protein, ARC, forms virus-like structures and is involved in transferring RNA between cells. Although these studies suggested that it might be possible to engineer retrotransposon proteins as a delivery platform, scientists had not successfully harnessed these proteins to package and deliver specific RNA cargoes in mammalian cells.
SEND packages are introduced to diseased cells to deliver therapeutic mRNA and restore health. Credit: McGovern Institute
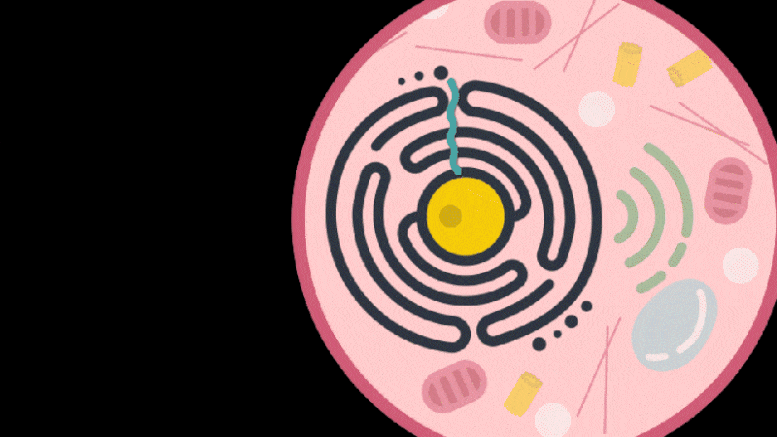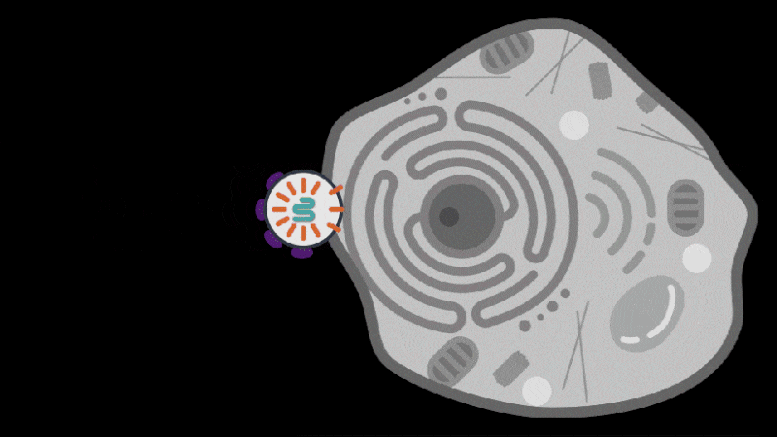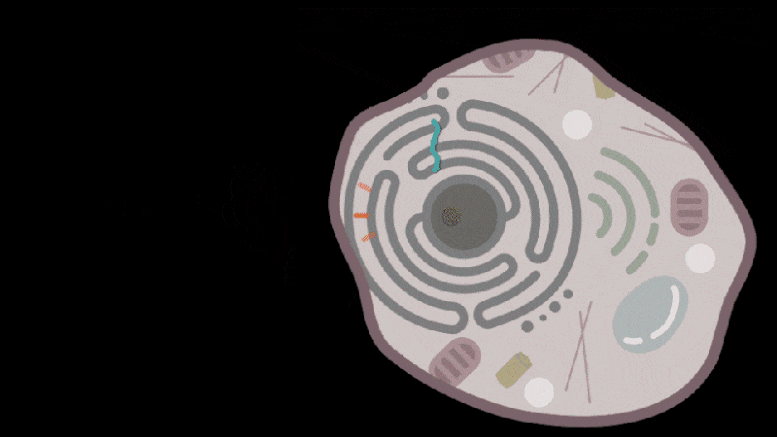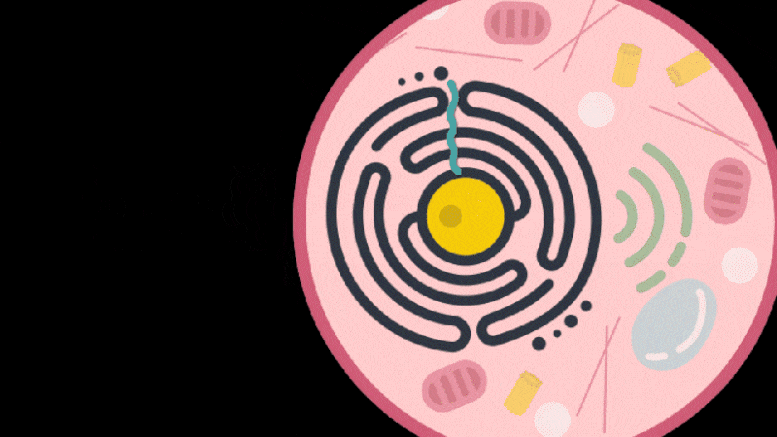
Knowing that some retrotransposon-derived proteins are able to bind and package molecular cargo, Zhang’s team turned to these proteins as possible delivery vehicles. They systematically searched through these proteins in the human genome for ones that could form protective capsules. In their initial analysis, the team found 48 human genes encoding proteins that might have that ability. Of these, 19 candidate proteins were present in both mice and humans. In the cell line the team studied, PEG10 stood out as an efficient shuttle; the cells released significantly more PEG10 particles than any other protein tested. The PEG10 particles also mostly contained their own mRNA, suggesting that PEG10 might be able to package specific RNA molecules.
Developing a modular system
To develop the SEND technology, the team identified the molecular sequences, or “signals,” in PEG10’s mRNA that PEG10 recognizes and uses to package its mRNA. The researchers then used these signals to engineer both PEG10 and other RNA cargo so that PEG10 could selectively package those RNAs. Next, the team decorated the PEG10 capsules with additional proteins, called “fusogens,” that are found on the surface of cells and help them fuse together.
By engineering the fusogens on the PEG10 capsules, researchers should be able to target the capsule to a particular kind of cell, tissue, or organ. As a first step towards this goal, the team used two different fusogens, including one found in the human body, to enable delivery of SEND cargo.
“By mixing and matching different components in the SEND system, we believe that it will provide a modular platform for developing therapeutics for different diseases,” said Zhang.
Advancing gene therapy
SEND is composed of proteins that are produced naturally in the body, which means it may not trigger an immune response. If this is demonstrated in further studies, the researchers say SEND could open up opportunities to deliver gene therapies repeatedly with minimal side effects. “The SEND technology will complement viral delivery vectors and lipid nanoparticles to further expand the toolbox of ways to deliver gene and editing therapies to cells,” said Lash.
Next, the team will test SEND in animals and further engineer the system to deliver cargo to a variety of tissues and cells. They will also continue to probe the natural diversity of these systems in the human body to identify other components that can be added to the SEND platform.
“We’re excited to keep pushing this approach forward,” said Zhang. “The realization that we can use PEG10, and most likely other proteins, to engineer a delivery pathway in the human body to package and deliver new RNA and other potential therapies is a really powerful concept.”
Reference: “Mammalian retrovirus-like protein PEG10 packages its own mRNA and can be pseudotyped for mRNA delivery” by Michael Segel, Blake Lash, Jingwei Song, Alim Ladha, Catherine C. Liu, Xin Jin, Sergei L. Mekhedov, Rhiannon K. Macrae, Eugene V. Koonin and Feng Zhang, 20 August 2021, Science.DOI: 10.1126/science.abg6155
This work was made possible with support from the Simons Center for the Social Brain at MIT; National Institutes of Health Intramural Research Program; National Institutes of Health grants 1R01-HG009761 and 1DP1-HL141201; Howard Hughes Medical Institute; Open Philanthropy; G. Harold and Leila Y. Mathers Charitable Foundation; Edward Mallinckrodt, Jr. Foundation; Poitras Center for Psychiatric Disorders Research at MIT; Hock E. Tan and K. Lisa Yang Center for Autism Research at MIT; Yang-Tan Center for Molecular Therapeutics at MIT; Lisa Yang; Phillips family; R. Metcalfe; and J. and P. Poitras.
SARS-CoV-2 uses its spike protein to attatch to a host cell.
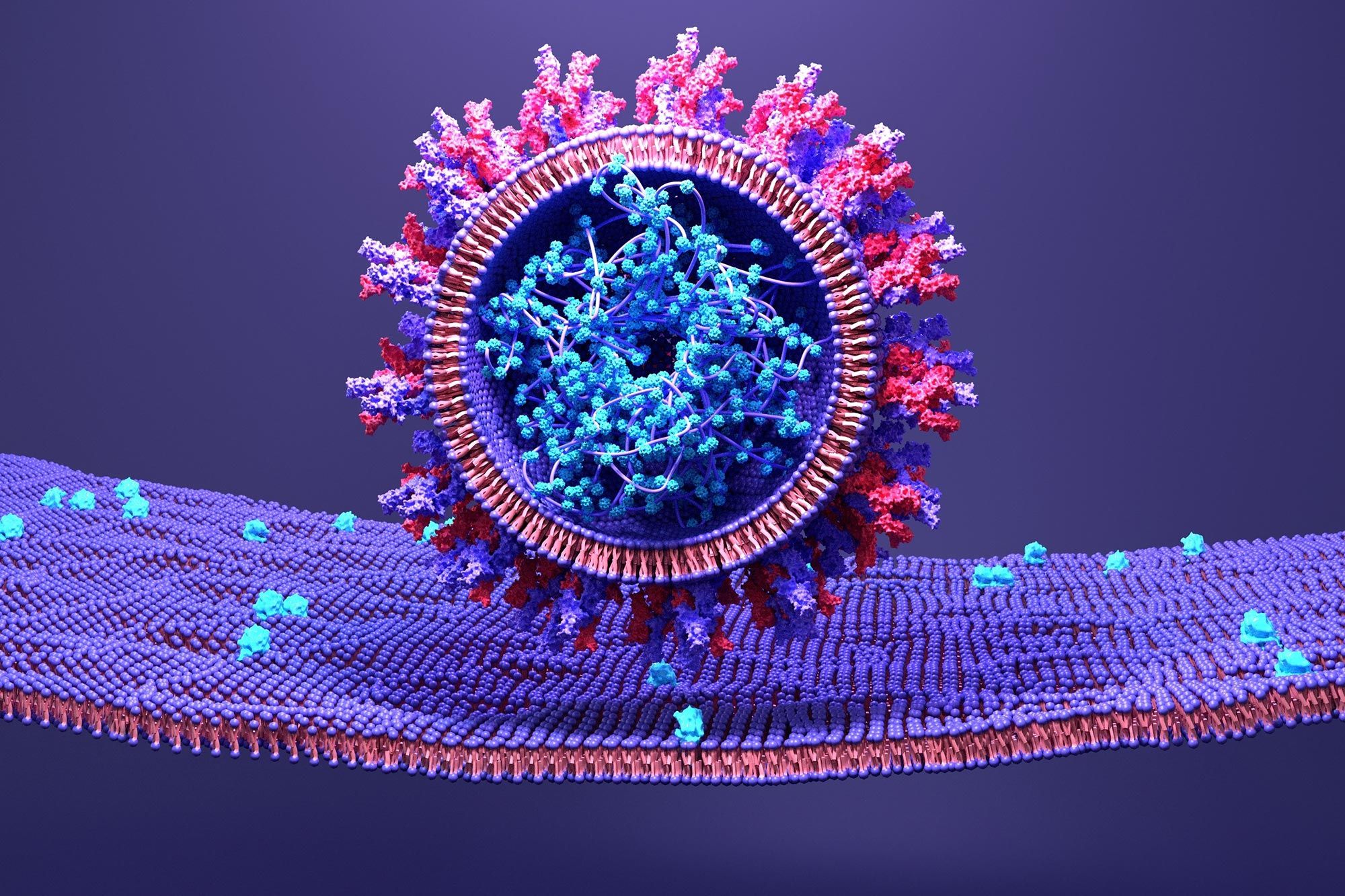
Therapeutic approach developed by Weizmann Institute scientists could spell new hope in the battle against COVID-19.
Even though vaccines may be steering the world toward a post-pandemic normal, a constantly mutating SARS-CoV-2 necessitates the development of effective drugs. In a new study published in Nature Microbiology, Weizmann Institute of Science researchers, together with collaborators from the Pasteur Institute, France, and the National Institutes of Health (NIH), USA, offer a novel therapeutic approach to combating the notorious virus. Rather than targeting the viral protein responsible for the virus entering the cell, the team of researchers addressed the protein on our cells’ membrane that enables this entry. Using an advanced artificial evolution method that they developed, the researchers generated a molecular “super cork” that physically jams this “entry port,” thus preventing the virus from attaching itself to the cell and entering it.
Prof. Gideon Schreiber. A new approach for treating COVID-19 developed by means of artificial evolution. Credit: Weizmann Institute of Science

Most potential therapies (and present vaccines) for SARS-CoV-2 target the so-called “spike protein” found on the virus’s outer envelope. This protein, however, is prone to mutations that erode the efficacy of these treatments. “Since the virus is constantly evolving we have, instead, focused on the non-evolving human receptor called ACE2 that acts as the entry site for the virus,” says Prof. Gideon Schreiber of Weizmann’s Biomolecular Sciences Department, who supervised the new study. This approach is not susceptible to new emerging virus variants, which is one of the main challenges in fighting the pandemic.
ACE2, attached to the membrane of lung epithelial cells and other tissues, is an enzyme important for regulating blood pressure. Therefore, as tempting as it may be to simply block this receptor to prevent the entry of SARS-CoV-2, any such strategy must not interfere with ACE2’s function. Schreiber, whose lab specializes in studying interactions between proteins, set out to develop a small protein molecule that could bind to ACE2 better than SARS-CoV-2 does but without affecting the receptor’s enzymatic activity.
The researchers, led by Dr. Jiří Zahradník, a postdoctoral fellow in Schreiber’s group, began by identifying SARS-CoV-2’s binding domain: the relatively short sequence of building blocks within the larger spike protein that physically binds to ACE2. Using the virus’s own receptor binding domain as a weapon against it, Zahradník performed several rounds of “evolution-in-a-test-tube,” developed in Schreiber’s lab, on a genetically engineered strain of baker’s yeast. Since yeast can be easily manipulated, Zahradník was able to rapidly scan millions of different mutations that accumulated in the course of this artificial evolution, a process that imitates natural evolution at a much faster pace. Ultimately, the goal was to find a small molecule that would be significantly “stickier” than the original viral version.
The rapid evolutionary process resulted in a small protein fragment with a binding capability 1,000 times stronger than that of the original viral binding domain
During this scanning process, Schreiber’s team supplied strong evidence in favor of the hypothesis that SARS-CoV-2 becomes more contagious when mutations improve its fit to ACE2. The researchers found that already after the first round of selection, the lab-made variants with tighter binding capabilities to ACE2 mimicked the mutations present in the binding domains of the most contagious SARS-CoV-2 strains that had occurred by means of natural evolution, such as the British variant (Alpha), the South African variant (Beta) and the Brazilian variant (Gamma). Surprisingly, the now widespread Indian (Delta) variant relies on a different trick to be more infectious – by partially evading detection by the immune system.
Structure of an ACE2 receptor (left), the original binding molecule (top right) and the newly engineered “super cork” (bottom right), imaged by cryogenic electron microscopy performed by Staff Scientists Dr. Nadav Elad of Weizmann’s Chemical Research Support Department and Dr. Orly Dym of the Life Sciences Core Facilities Department. The black outline indicates the binding site of the “super cork” on the ACE2 receptor. Credit: Weizmann Institute of Science
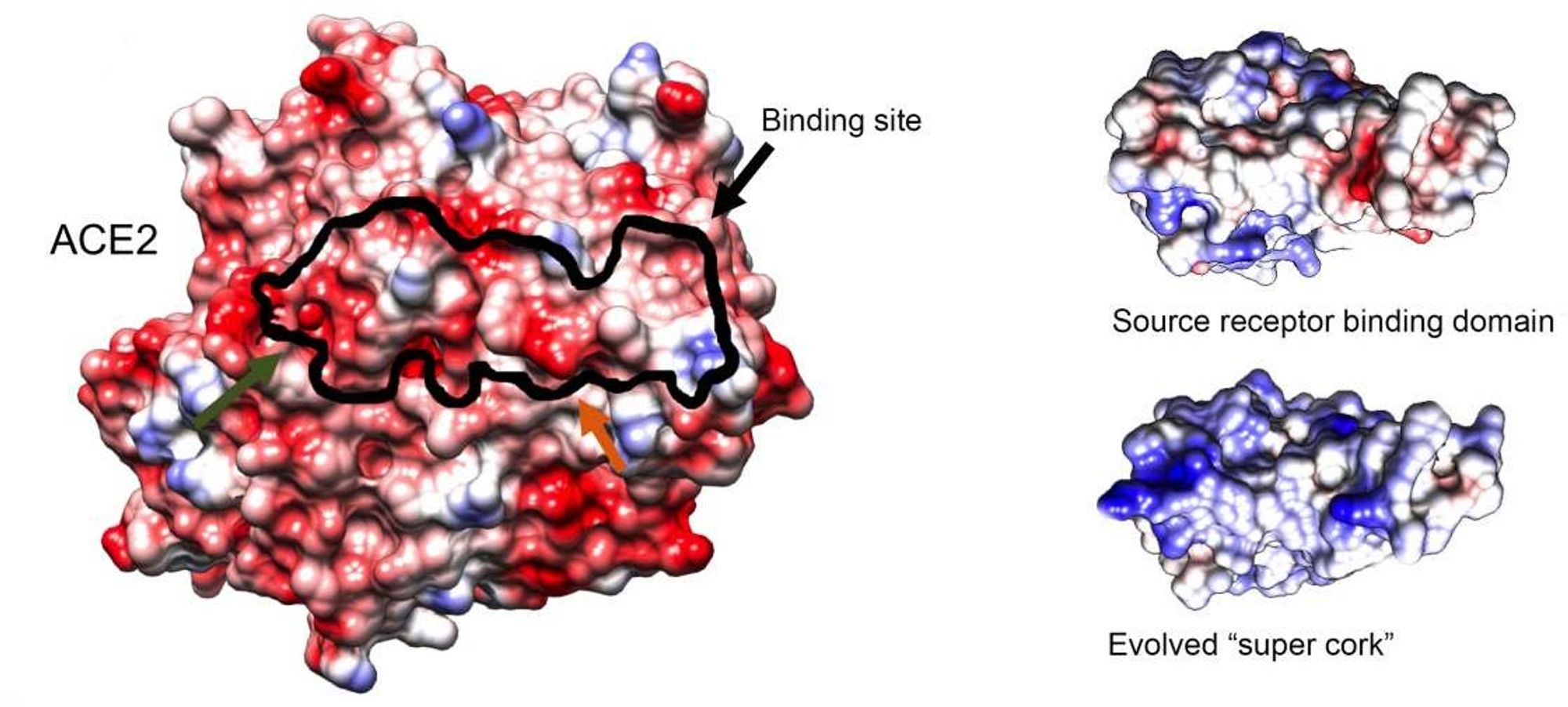
Eventually, Zahradník isolated a small protein fragment with a binding capability 1,000 times stronger than that of the original binding domain from which it evolved. This “super cork” not only fit ACE2 like a glove, it was also found by Maya Shemesh and Shir Marciano, PhD students in Schreiber’s lab, to conserve the enzymatic activity of ACE2 – just as the researchers had intended. Furthermore, owing to the strong binding, very low concentrations of the newly engineered molecule were needed to achieve the desired blocking effect.
While searching for a “super cork” that would block the ACE2 receptor, the researchers examined about 1,000,000,000 yeast mutants.
To develop a potential method for administering the molecule as a drug, Schreiber and his team collaborated with Prof. Yinon Rudich of Weizmann’s Earth and Planetary Sciences Department. Together with Dr. Ira Marton and Dr. Chunlin Li, they created an aerosol-based spray that would allow the developed molecule to be administrated by inhalation to patients.
So far, the researchers have tested, at the NIH, the developed formulation in hamsters infected with SARS-CoV-2. Preliminary results indicate that this treatment significantly reduces disease symptoms, suggesting that it could be a potential drug. More preclinical studies are planned to take place at the NIH in the near future.
Reference: “SARS-CoV-2 variant prediction and antiviral drug design are enabled by RBD in vitro evolution” by Jiří Zahradník, Shir Marciano, Maya Shemesh, Eyal Zoler, Daniel Harari, Jeanne Chiaravalli, Björn Meyer, Yinon Rudich, Chunlin Li, Ira Marton, Orly Dym, Nadav Elad, Mark G. Lewis, Hanne Andersen, Matthew Gagne, Robert A. Seder, Daniel C. Douek and Gideon Schreiber, 16 August 2021, Nature Microbiology.DOI: 10.1038/s41564-021-00954-4
Prof. Gideon Schreiber’s research is supported by the Ben B. and Joyce E. Eisenberg Foundation; the Rene and Tillie Molho Family Trust; Miel de Botton; and the Yotam Project.
Phobos imaged by HiRISE. Credit: NASA/JPL-Caltech/University of Arizona
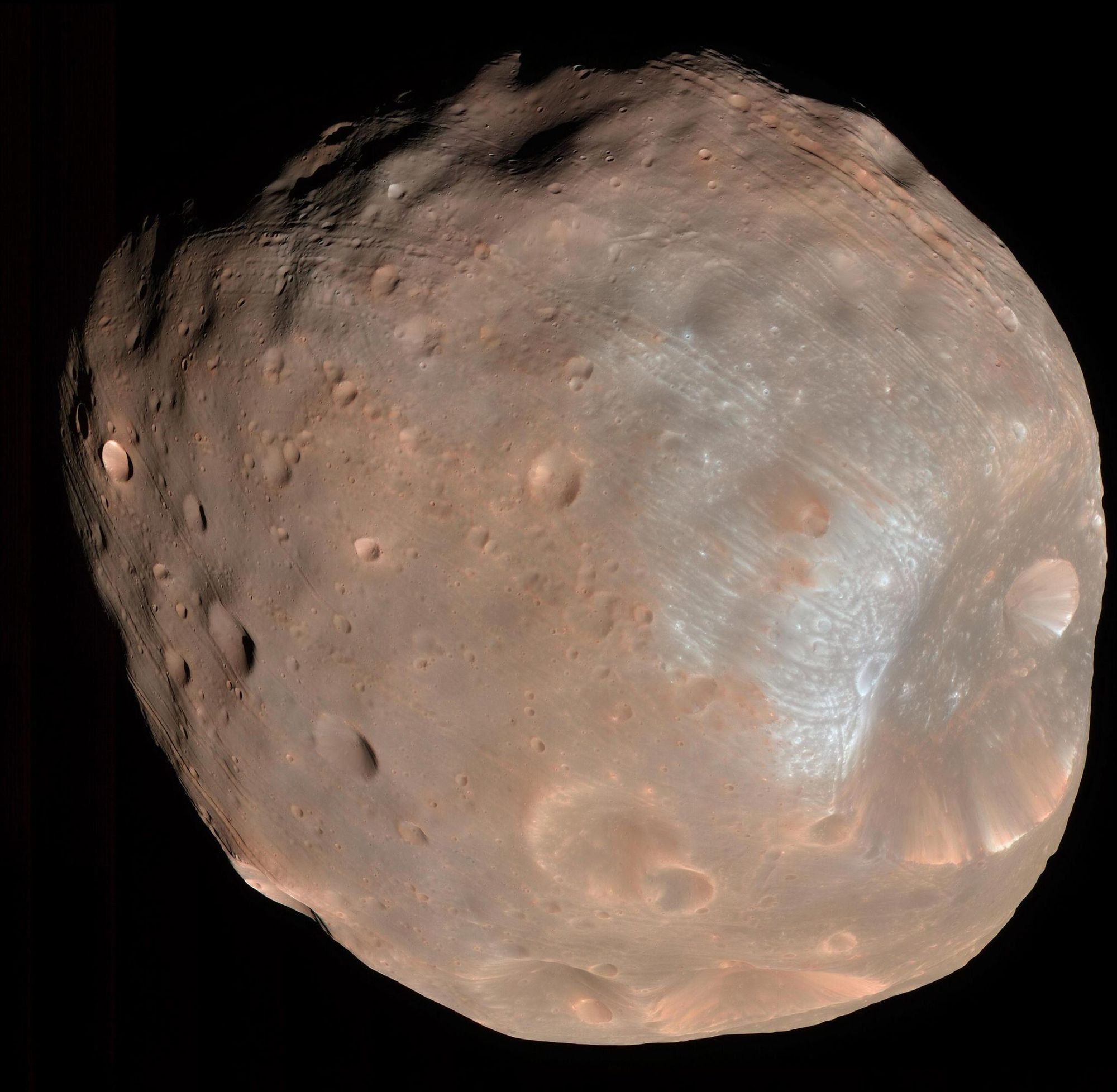

Japan’s space agency (JAXA) is gearing up for its Martian Moons eXploration (MMX) mission, with plans to have a sample from Mars’ moon Phobos return to Earth by 2029. Mission scientists say they hope to find clues to the origins of Mars two moons, as well as Mars itself, and possibly even traces of past life.
“We think that the Martian moon, Phobos, is loaded with material lifted from Mars during meteorite impacts,” the MMX team said on Twitter. “By collecting this Phobos sample, MMX will help investigate traces of Martian life and the new era of Martian habitability exploration in the 2020s will begin.”
MMX currently has a planned launch in 2024, with the spacecraft reaching the Mars system by 2025, approximately one year after leaving Earth. Current plans for the mission include an orbiter, a lander that touches down on Phobos with sample collection and return capability, and perhaps even a rover.
A newly released overview of the MMX mission. Credit: JAXA
The orbiter will be placed in a so-called Quasi Satellite Orbit (QSO) around Phobos, to collect scientific data. After both orbital and in situ observations, and sample collection, the lander spacecraft will liftoff to return to Earth carrying the sample of material gathered from Phobos. Current plans are for the lander to collect 10 grams (0.35 ounce) of soil. In a news briefing this week, and reported by the Associated Press, JAXA scientists said that perhaps about 0.1% of the surface soil on Phobos came from Mars, and 10 grams could contain about 30 granules, depending on the consistency of the soil.
The team says their exploration of the Martian moons will help improve technology for future planet and satellite exploration. They say their mission will aid in advancements, for example, in the technology required to make round-trips between the Earth and Mars, enhanced sampling techniques and optimal communication technology using the Deep Space Network ground stations.
The objectives of the MMX mission are:
- To investigate whether the Martian moons, Phobos and Deimos, are captured asteroids or fragments that coalesced after a giant impact with Mars, and to acquire new knowledge on the formation process of Mars and the terrestrial planets.
- To clarify the mechanisms controlling the surface evolution of the Martian moons and Mars, and to gain new insights into the history of the Mars Sphere, including that of the Martian moons.
MMX Infographic. Credit: JAXA/NASA
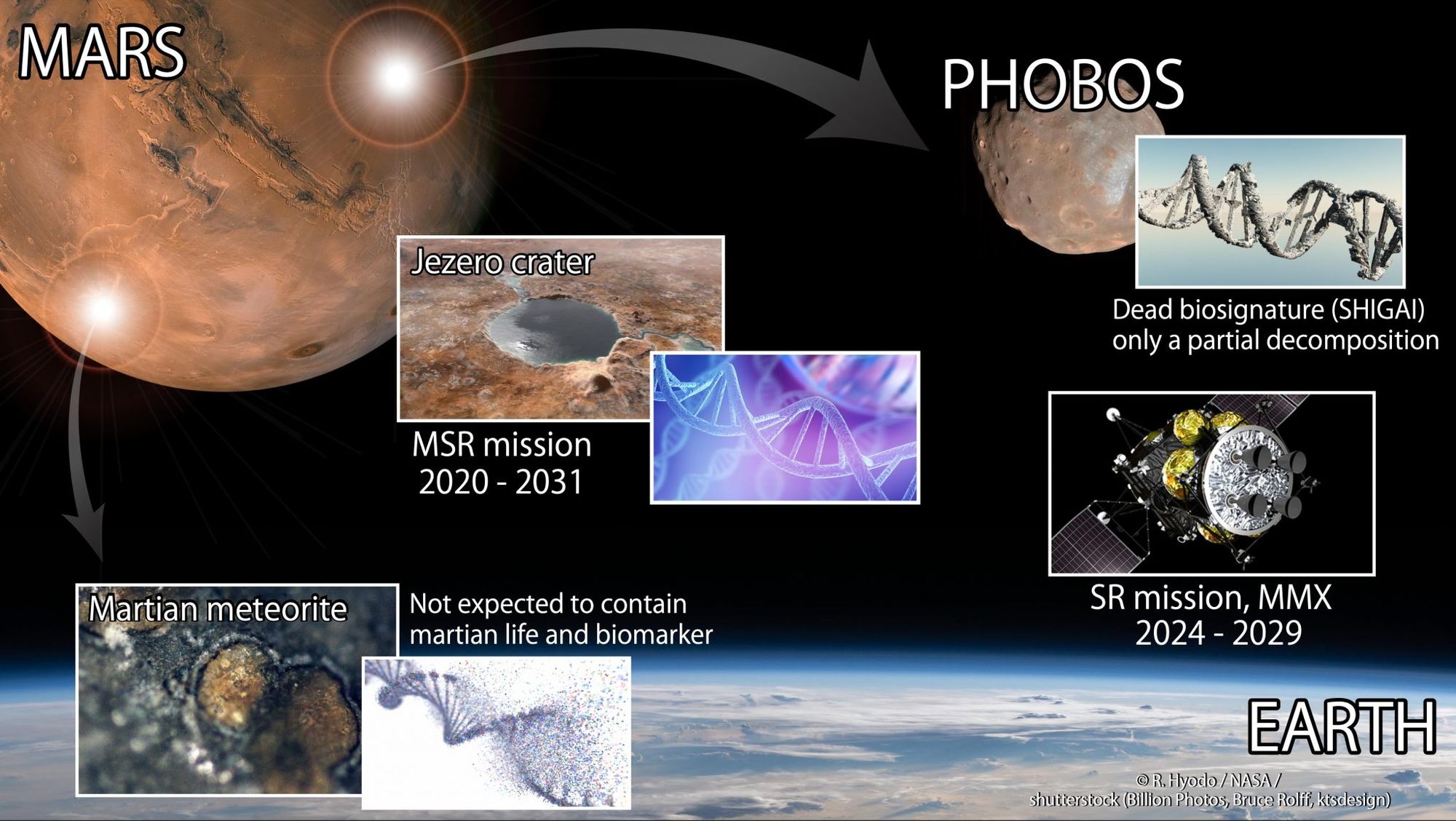

Sending a mission to moons of Mars has long been on the wish list for mission planners and space enthusiasts, and for the past few years, JAXA engineers and scientists have been working on putting such a mission together.
Many scientists say that studying and landing on the moons of Mars would be the next best thing to going to Mars itself. Phobos and Deimos have been considered as places for a possible human base that would allow for easier access to Mars than going to the Red Planet directly, especially for the first human missions to the Mars system.
“Humans can realistically explore the surfaces of only a few objects and Phobos and Deimos are on that list,” NASA Chief Scientist, Jim Green said in 2020. “Their position orbiting about Mars may make them a prime target for humans to visit first before reaching the surface of the Red Planet, but that will only be possible after the results of the MMX mission have been completed.”
Artist’s concept of the MMX spacecraft in orbital configuration, with its scientific instruments indicated. Credit: JAXA/ISAS
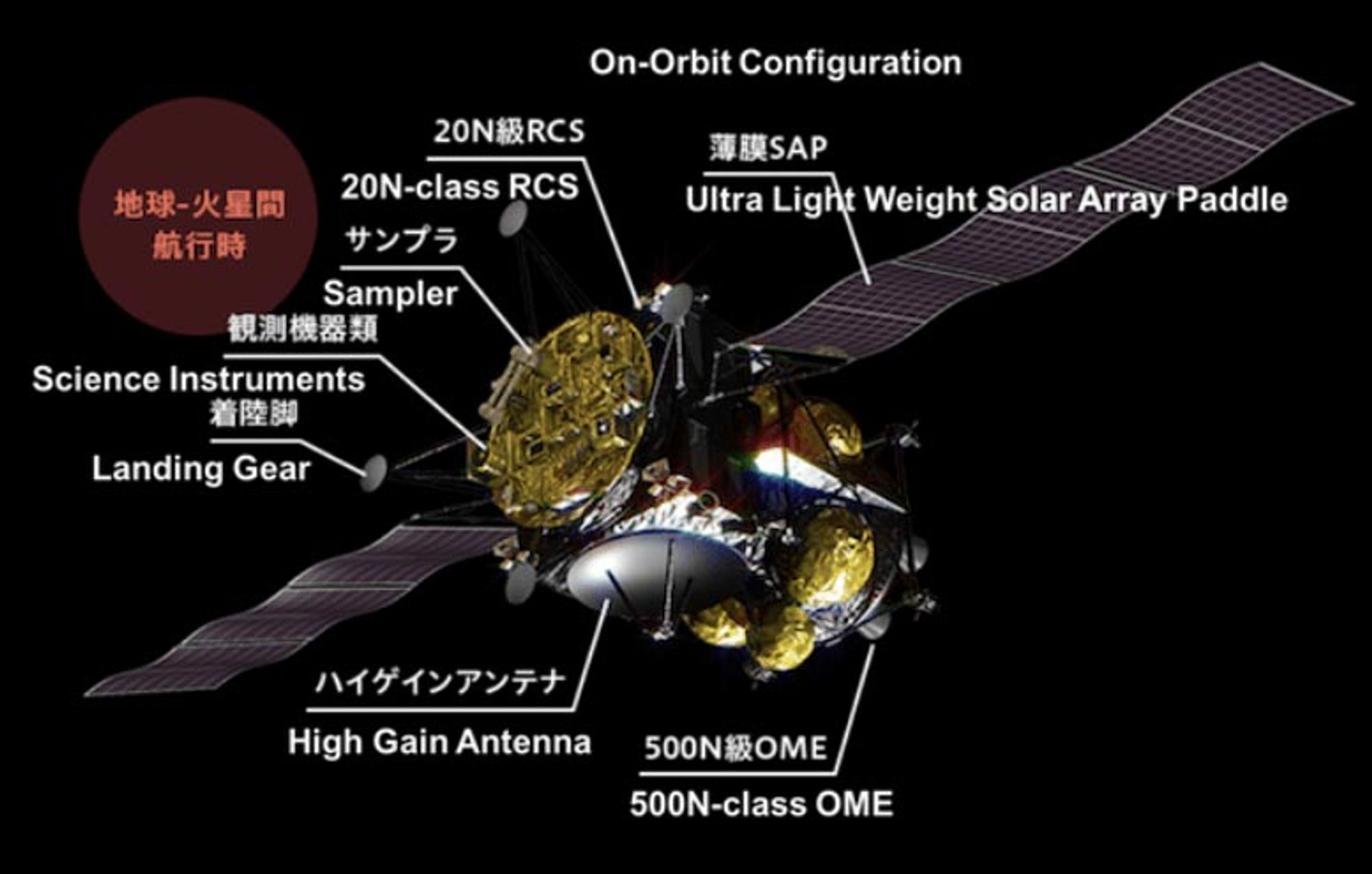
The mission will have international input, equipped with eleven instruments, four of which will be provided by international partners at NASA (USA), ESA (Europe), CNES (France) and DLR (Germany).
The JAXA-built instruments include a telescopic (narrow-angle) camera for observing detailed terrain, the wide-angle camera for identifying hydrated minerals and organic matter, the LIDAR laser altimeter, a dust monitor, and a mass spectrum analyzer, to study the charged ions around the moons, the sampling device and sample return capsule, and a radiation environment monitor.
NASA has signed on to contribute a gamma ray and neutron spectrometer to examine the elements that constitute the Martian moons, and also a pneumatic sampling device. CNES is building a near-infrared spectrometer that can identify mineral composition, and is working with DLR to design the rover, which could explore the surface of Phobos. ESA is listed as assisting with deep space communication equipment.
The quick turnaround time for MMX’s sample return would put Japan ahead of the United States and China in bringing back samples from the Mars system, even though they started later, said MMX project manager Yasuhiro Kawakatsu in this week’s news conference.
A new paper published by JAXA’s Institute of Space and Astronautical Research proposes that if Mars was once a habitable planet, with water on the surface, the water could have been ejected into space by gigantic Martian dust storms. They say that material collected by MMX from Phobos will help confirm if this theory might be true.
Originally published on Universe Today.