Listen to this story
Google researchers recently announced Segmentation-Guided Contrastive Learning of Representations (SegCLR), a new model that can produce a high-resolution 3D mapping of brain cells and their connectivities.
This method can train rich, generic representations of cell shape and internal structure effortlessly without manual labelling, producing compact vector representations (embeddings) applicable across diverse downstream tasks. Some examples include local classification of cellular subcompartments and unsupervised clustering. Additionally, the model can recognise different cell types, even from small fragments of a cell.
The research is an important breakthrough in neuroscientific research as the prediction model can lead to an understanding of how brain circuits develop and function in health and disease.
The researchers trained SegCLR on the H01 human cortex dataset and the MICrONS mouse cortex dataset. The resulting embedding vectors for both the datasets, about 8 billion in total, will be released for exploring further research.
Interestingly, their findings suggest that for identifying cellular subcompartments (axon, dendrite, soma, etc.), a straightforward linear classifier trained on top of SegCLR embeddings outperformed a fully supervised deep network trained on the same task, with only one thousand labelled instances instead of millions.
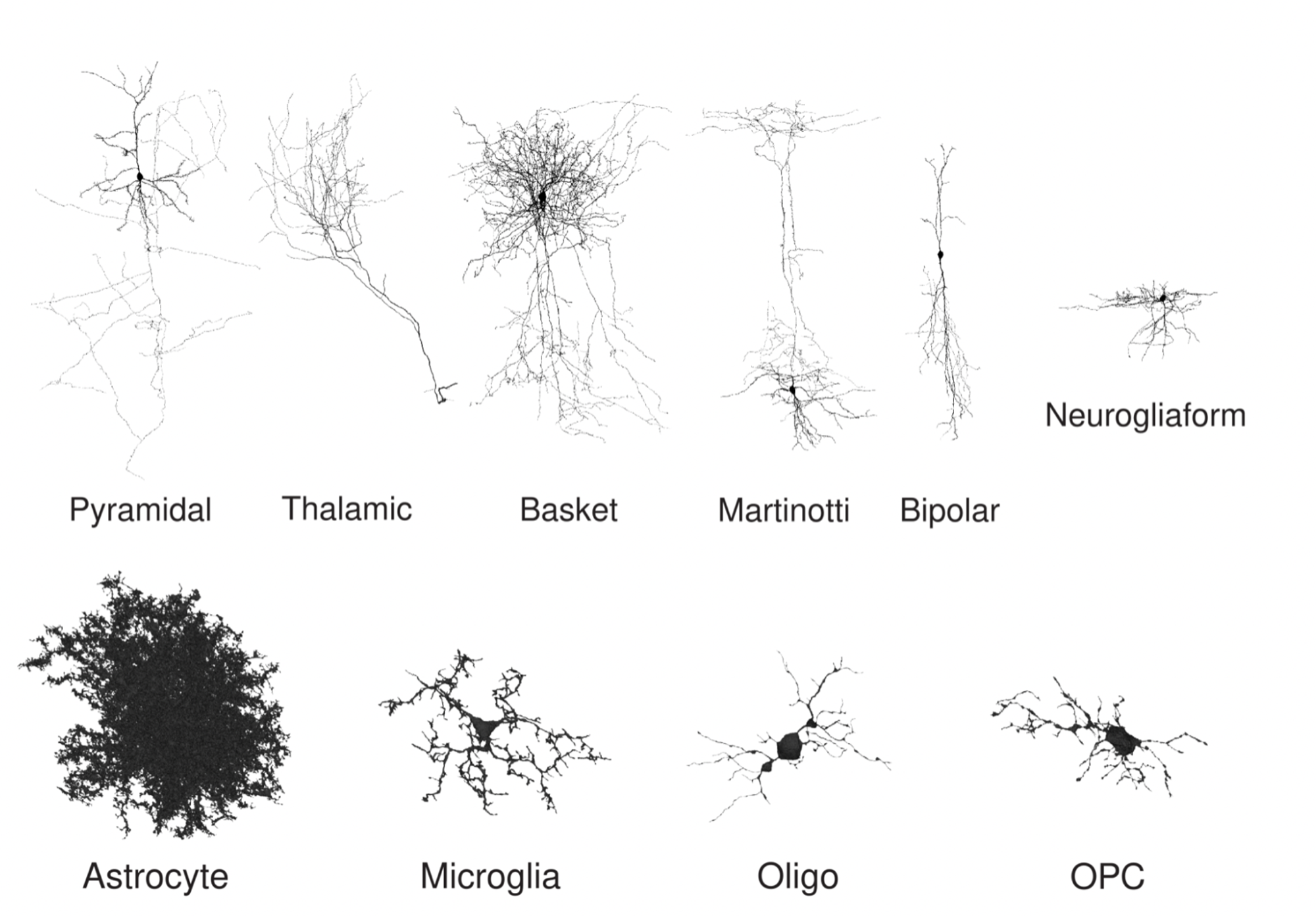
Before classifying the data, the researchers gathered and averaged embeddings within each cell over a set aggregation distance, which is the distance from a central point. The model predicted human cortical cell types with high accuracy, even for aggregation radii as small as 10 micrometres. This includes for types that even experts find difficult to distinguish, such as microglia (MGC) versus oligodendrocyte precursor cells (OPC).